This Evolution 101 post is by MSU grad student Douglas Kirkpatrick

Fig. 1: Lebron James helps lead the Cavaliers in a sweep of the Celtics.
In baseball, ice hockey, and basketball, when a team wins all of the games in a playoff series, they are said to have swept the other team out of the playoffs. In other words, a sweep is the complete victory of one group over the other, often due to a star player. Funnily enough, this terminology applies almost as well in the world of evolutionary biology. A selective sweep is the process by which strong selection of a beneficial allele causes the surrounding linked alleles to achieve a higher frequency in the population. In the same way that superstar Lebron James lead his team, the Cleveland Cavaliers, to a sweep of the Hawks and the Celtics in last year’s playoffs, a good gene might help linked alleles attain reproductive success in a selective sweep.
The essence of this process is that when a hugely beneficial allele appears, it is rapidly selected for. Due to the rapid selection, many linked or nearby genes are also passed on to descendants, even though these other alleles may be neutral or even mildly deleterious to the organism. This process is illustrated in Figure 2. The extremely beneficial gene is highlighted in red, while deleterious or neutral genes are presented in blue. Each line in Figure 2 represents the genome of a single member of the population. The pattern of the beneficial gene and its neighbors, seen in the fourth row on the right, are collectively referred to as a “haplotype.” The haplotype of the advantageous gene becomes much more frequent after selection occurs, as new populations are spawned [3].
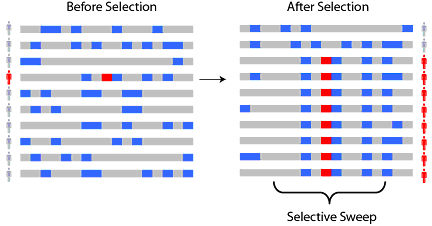
Fig 2: A basic biological selective sweep
Several methods exist to detect selective sweeps; the primary method is to measure linkage disequilibrium. That is, the distributions of alleles in a population are compared to determine the presence of haplotypes. If a specific haplotype, or collection of linked alleles, is exceedingly common, then it is likely that a selective sweep occurred in the recent past. The linked alleles are primarily those that are collocated within the genome. An alternative measure to find instances of selective sweep is to measure the time to most recent common ancestor. If they have all evolved from a recent ancestor, then it is likely that a selective sweep has occurred. That is, because the ancestor is recent, its haplotypes have likely spread quickly through the population, making it probable that a selective sweep has occurred.
Two examples of selective sweeps are particularly relevant in the modern world: in pathogenic bacteria and in agriculture. Bacteria and disease-causing bacteria in particular, have a short life span. As such, any new allele that creates a more virulent form will spread wildly as the less potent haplotypes die off in a short time. The potential for rapid change in addition to strong selective pressure from outside forces like antibiotics or antivirals leads to many selective sweeps. The short life cycles and high selective pressures placed on pathogens have been shown to have caused selective sweeps in influenza and toxoplasma gondii [1].
While the selective sweeps in bacteria may be detrimental to mankind, there are selective sweeps that have helped humanity. Data shows that selective sweeps were responsible for unifying a diverse population into what we know as modern corn. Artificial pressure, driven by farmers only choosing optimal offspring, in coordination with selective crossbreeding forced a fast evolution of corn. The optimal traits selected for and the strong evolutionary pressure created a selective sweep [5]. The selective sweep in this case proved to be largely beneficial for people.
Figure 3: Selective Sweeps in human populations. Tishkoff et. al, 2007
Numerous examples of selective sweeps can be found in human DNA. There are at least six different chromosomes that show evidence of selective sweeps [4]. The most noticeable was described by Tishkoff et al., and comes as a result of the alleles for lactose tolerance. There was strong selection for the alleles that allow humans to digest milk as adults. As a result, selective sweeps occurred; this is illustrated in Figure 3. Interestingly, similar mutations occurred separately in Africa (Group A) and Eurasia (Group B). Each individual bar represents a portion of an individual’s genome that is shared with the group; a larger bar indicates that more genetic material is the same as the original cell. The horizontal axis is the relative position to the allele for lactase tolerance. The red and the green in each graph respectively show the haplotype for tolerance that has become more common, while the blue and orange are an old haplotype that has been outcompeted [2]. This information is essential to understanding how different yet similar processes evolved in humans; it is likely to be important going forward due to how many selective sweeps have occurred in the human genome.
Selective sweeps are important because they allow for rapid evolution in a short period of time. The strong selection for a specific haplotype can quickly change the distribution of alleles in a population. In addition, finding a selective sweep can help identify key periods of evolutionary change. Sweeps have had a major impact both on the human genome, and that of plants and animals seen in everyday life; they will continue to do so into the foreseeable future.
Sources:
- Sa, Juliana Marth, et al. (2009). “Geographic patterns of Plasmodium falciparum drug resistance distinguished by differential responses to amodiaquine and chloroquine”. PNAS 106 (45): 18883–18889.
- Tishkoff, Sarah A., et al. “Convergent adaptation of human lactase persistence in Africa and Europe.”Nature genetics 1 (2007): 31-40.
- A selective sweep. http://www.nature.com/scitable/content/a-selective-sweep-24827
- “A haplotype map of the human genome”. Nature 437 (7063): 1299–1320. October 2005. doi:10.1038/nature04226
- Gore, Michael A., et al. (2009). “A First-Generation Haplotype Map of Maize”. Science 326: 1115–1117. doi:10.1126/science.1177837. PMID 19965431.
